STRING API
STRING has an application programming interface (API) which enables you to get the data without using the graphical user interface of the web page. The API is convenient if you need to programmatically access some information but still do not want to download the entire dataset. There are several scenarios when it is practical to use it. For example, you might need to access some interaction from your own scripts or want to incorporate some information in STRING to a web page.
We currently provide an implementation using HTTP, where the database information is accessed by HTTP requests. Due to implementation and licensing reasons, The API provide methods to query individual items only, similar to the web site. If you need access to bulk data, you can download the entire dataset by signing the academic license agreement.
API format Top ↑
To call the API, make a URI (uniform resource identifier) of the following form:
http://[database]/[access]/[format]/[request]?[parameter]=[value]
Possible values for the fields are listed in the tables below. Note that only some of the combinations of the fields are possible as shown on this diagram:
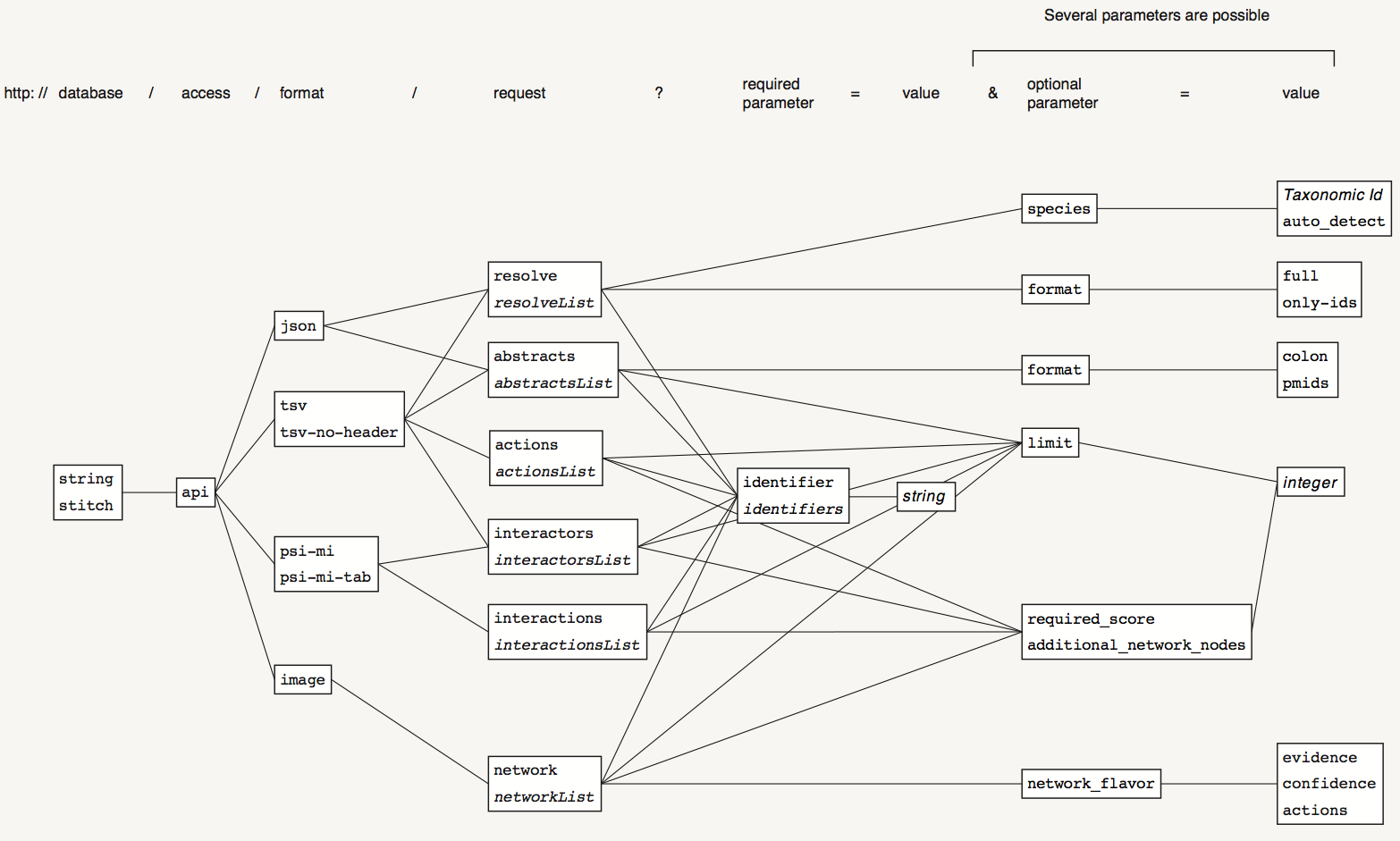
Download PDF (1073x645 pixels, file size: 119 KB)
As an identifier, it is possible to use either a
single entry or a list of entries. The parameter is then called
identifiers or identifiers,
respectively. In the former case, the
request should be
resolve,
abstracts,
actions,
interactors,
interactions or network
followed by the required parameter identifier
and its value. In the latter case, when there are several entries, the
request should be ending with
...List and have the required parameter
identifiers.
The values of the identifiers parameter contains a list of items
separated by new line characters (%0A or %0D).
It is also possible to use a single item in list of identifies, for most cases this will be identical to the single item queries mentioned above.
List of databases Top ↑
| Database | Description |
|---|---|
| string-db.org | Main entry point of STRING |
| string.embl.de | Alternative entry point of STRING |
| stitch.embl.de | The sister database of STRING |
List of accesses Top ↑
| Access | Description |
|---|---|
| api | Application programming interface |
| services | Other services to access data |
List of return formats Top ↑
| Format | Description |
|---|---|
| json | JSON format either as a list of hashes/dictionaries, or as a plain list (if there is only one value to be returned per record) |
| tsv | Tab separated values, with a header line |
| tsv-no-header | Tab separated values, without header line |
| psi-mi | The interaction network in PSI-MI 2.5 XML format |
| psi-mi-tab | Tab-delimited form of PSI-MI (similar to tsv, modeled after the IntAct specification. (Easier to parse, but contains less information than the XML format.) |
List of requests Top ↑
| Request | Returns |
|---|---|
| resolve | List of items that match (in name or identifier) the query item |
| resolveList | List of items that match (in name or identifier) the query items |
| abstracts | List of abstracts that contain the query item |
| abstractsList | List of abstracts that contain any of the query items |
| interactors | List of interaction partners for the query item |
| interactorsList | List of interaction partners for any of the query items |
| actions | Action partners for the query item |
| actionsList | Action partners for any of the query items |
| interactions | Interaction network in PSI-MI 2.5 format or PSI-MI-TAB format (similar to tsv) |
| interactionsList | Interaction network as above, but for a list of identifiers |
| network | The network image for the query item |
| networkList | The network image for the query items |
List of parameters and values Top ↑
| Parameter | Description |
|---|---|
| identifier | required parameter for single item, e.g. DRD1_HUMAN |
| identifiers | required parameter for multiple items, e.g. DRD1_HUMAN%0DDRD2_HUMAN |
| format | For resolve requests: only-ids get the list of only the STRING identifiers (full by default). For abstract requests: use colon pmids for alternative shapes of the pubmed identifier |
| species | Taxon identifiers (e.g. Human 9606, see: http://www.uniprot.org/taxonomy) |
| limit | Maximum number of nodes to return, e.g 10 |
| required_score | Threshold of significance to include a interaction, a number between 0 and 1000 |
| additional_network_nodes | Number of additional nodes in network (ordered by score), e.g. 10 |
| network_flavor | The style of edges in the network. evidence for colored multilines. Confidence for singled lines where hue correspond to confidence score. (actions for stitch only) |
| caller_identity | Your identifier for us. |
The meaning of the columns returned from STRING API calls Top ↑
| Column | Description |
|---|---|
| nscore | neighborhood score, (computed from the inter-gene nucleotide count) |
| fscore | fusion score (derived from fused proteins in other species) |
| pscore | cooccurence score of the phyletic profile (derived from similar absence/presence patterns of genes) |
| hscore | homology score, the degree of homology of the interactors (normally not reported in STRING) |
| ascore | coexpression score (derived from similar pattern of mRNA expression measured by DNA arrays and similar technologies) |
| escore | experimental score (derived from experimental data, e.g., affinity chromatography) |
| dscore | database score (derived from curated data of various databases) |
| tscore | textmining score (derived from the co-occurrence of gene/protein names in abstracts) |
Examples Top ↑
Find out which proteins match the description "ADD" in human:
http://string-db.org/api/tsv/resolve?identifier=ADD&species=9606
Return bare IDs that you could pipe into other STRING API-functions:
http://string-db.org/api/tsv-no-header/resolve?identifier=YOL086C&format=only-ids
Maximum 10 PMID (default setting for limit) for that contain any of that identifiers:
http://string-db.org/api/tsv/abstractsList?identifiers=4932.YML115C%0D4932.YJR075W%0D4932.YEL036C
Get the 20 highest scoring interactors above score 400 for a list of queries:
http://string-db.org/api/tsv-no-header/interactorsList?identifiers=4932.YML115C%0D4932.YJR075W%0D4932.YEL036C&required_score=400&limit=20
Gert interactions above score 900 for a list of queries and limit to 5 nodes: (i.e. max 5.6/2=15 interactions).
http://string-db.org/api/psi-mi-tab/interactionsList?identifiers=4932.YML115C%0D4932.YJR075W%0D4932.YEL036C&required_score=900
Retrieve an image of the network of a known identifier, with a score threshold of 950, limited to 10 interacting protein, and a evidence style of the edges:
http://string-db.org/api/image/network?identifier=4932.YML115C&required_score=950&limit=10&network_flavor=evidence
Additional information about the API Top ↑
As mentioned above, if you need to do large-scale analysis, please download full data set. Otherwise you may flood the string server with api requests, in particular if you run scripts in parallel.
The format of the API will stay constant with upcoming versions of STRING, but it is likely that add more query options later for retrieving information. For example: What are the synonyms of this item? To which orthologous group does this protein belong? etc.
The caller_identity parameter can be used for identifying you call to STRING. We can provide logging information according to such information. Please contact the string team if you have particular needs.